Biology tools for Rust
A coherent set of crates for file operations, MD, database/API integration, and numeric computation. These are the building blocks behind Molchanica and PlasCAD; you can embed them directly in your own software. Designed for structural biology and bioinformatics workflows, but some libraries have more general applications.
These are designed to simplify the creation and maintenance of biology and chemistry applications and libraries written in Rust.
Python bindings are available for most of these, installable with Pip or UV. Note that the Python ecosystem has existing high quality libraries for some of these uses.
Core libraries
Load and save mmCIF, Mol2, SDF, PDBQT, Map, Amber DAT/FRCMOD/lib, GRO, and more. Provides generic atom, bond, molecule, and residue structs you can use directly or convert into application-specific ones. Can load molecules remotely from databases (e.g. RCSB PDB, PubChem, DrugBank). Provides a high-level interface for ORCA. (Quantum chemistry)
High-level access to RCSB, PubChem , PDBe, DrugBank, NCBI BLAST, and Amber Geostd. Fetch structures, density, moedule files, metadata, perform queries, or open the corresponding browser pages.
Simulation tools and visualization
Runs classical forcefield-based MD with Amber force fields. For Rust and Python. Includes Velocity-Verlet and Langevin/CSVR integrators, thermostat and stochastic cell rescaling barostat, explicit OPC water, and the ability to load or compute force field parameters for arbitrary molecules. Run MD on any set of protein, nucleic acid, lipid, or small organic molecules. Computes force field parameters and partial charge for arbitrary molecules.
Easy to use, and performance-oriented: Uses GPU via CUDA, our CPU SIMD and thread pools for parallelization. Ideal for embedding MD in custom tools and applications.
Numerics and visualization
Calculate Smooth Particle Mesh Ewald (SPME) force. Fast coulomb forces with periodic boundary conditions. Uses CPU, or GPU via cuFFT or VkFFT (CUDA backend). Used by our MD tools to get order N log(N) scaling for long-range forces.
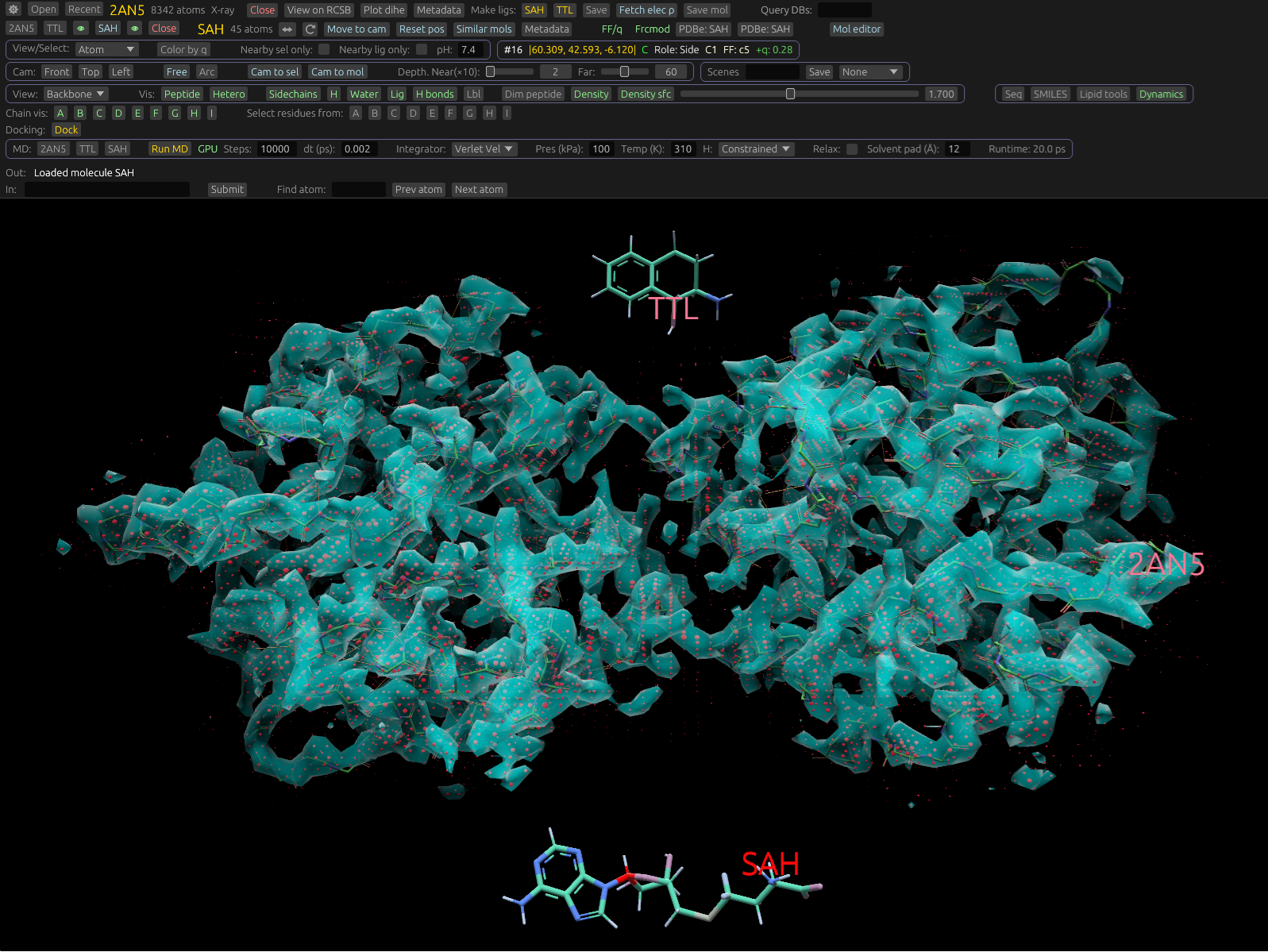
Marching Cubes for Rust/Python. Generates meshes from amplitudes. For example, this can be used to visualize electron density, and solvent-accessible surfaces. Used to render Cryo-EM and crystallography maps in molchanica. Outputs a simple mesh you can plug into your renderer. You can also compute the volume of the mesh.