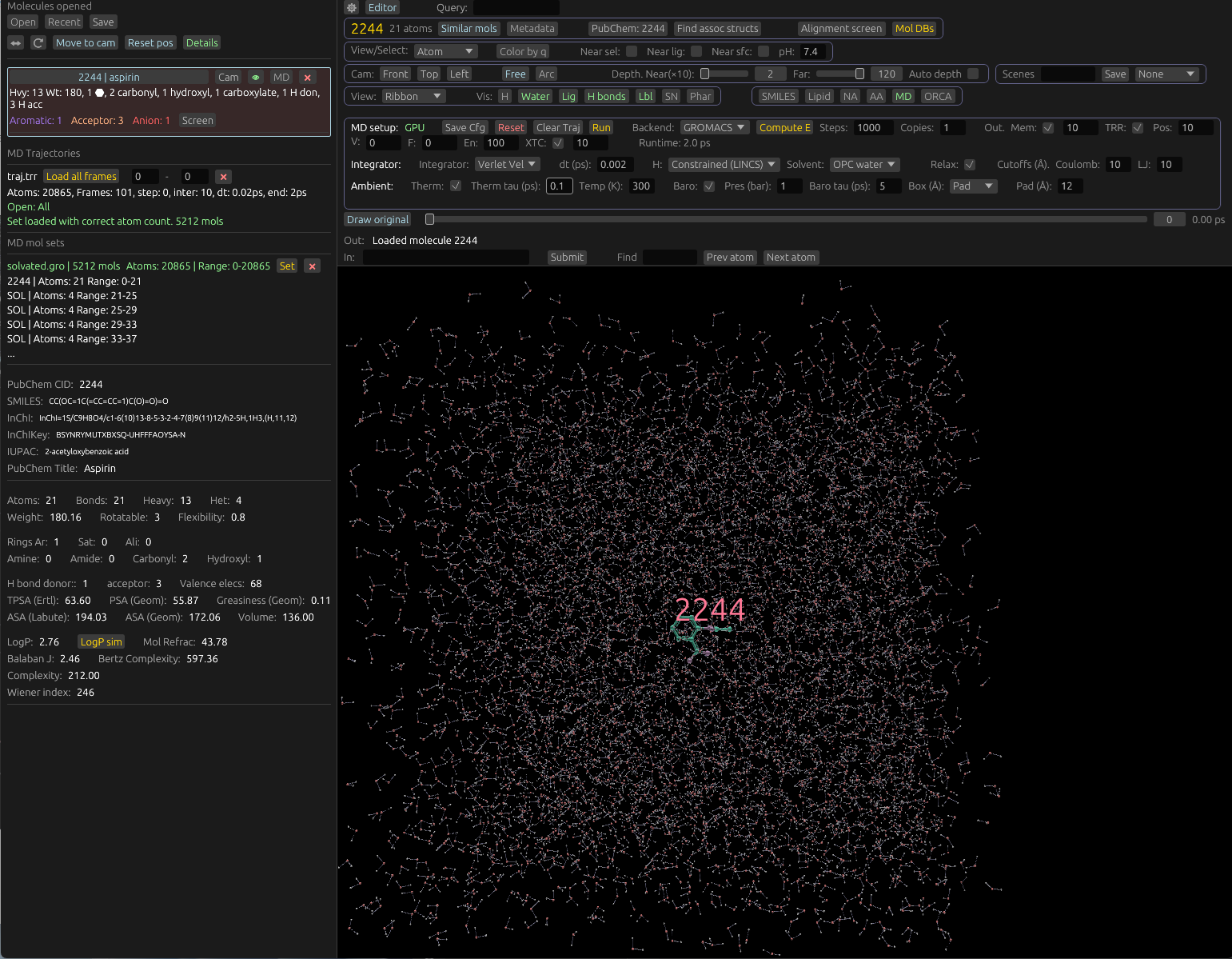
Molecular dynamics viewer / playback
Molchanica can play back MD trajectories generated by itself, and other programs. If running MD locally and memory is selected as an output option (Or if GROMACS is the backend and TRR is selected), frames will be automatically loaded into the viewer upon completion. When loading a trajectory, two components are required: The trajectory itself, and a molecule set.
Trajectories include frames, describing atom positions and velocities. These are listed by atom in a flattened structure, and don't delineate by molecule. Trajectories can be loaded in the following formats:
- TRR (GROMACS)
- XTC (GROMACS, compressed)
- DCD (NAMD, OpenMM, Amber, CHARMM)
Molecule sets define how we divide trajectories into molecules, for the purposes of visualization. Molecule sets are created upon an MD run in Molchanica, and are output as files by other programs. These formats have much in common with standard molecule formats (mmCIF, SDF, Mol2, etc), but usually contain multiple molecules.
Trajectories and molecule sets can be loaded using the same system as other files: Drag them into the window, or use the Open button. They will be displayed in the sidebar. If there is no molecule set loaded, trajectories will be marked with "No mol set loaded". If there is a molecule set loaded, but its atom count does not match the trajectory's, then a message will be displayed indicating this.
The main trajectory viewer controls will display a playback slider if ready. If not, it will show a message explaining way. For example, no trajectory loaded, no trajectory from snapshots, mismatch in MD set atom count, etc.
You can load all frames from a trajectory, or a subset of frames, using the associated entry in the sidebar. You can clear the trajectory frames from the viewer UI.
Click or drag the slider to change the active frame.
Creating and editing molecule sets
Molecule sets are automatically created and loaded after running MD through Molchanica. If using the GROMACS backend, they're saved as .gro files automatically in the GROMACS output directory. (e.g. gromacs_out). To create or edit a new set, click "Edit mol sets" in the MD sets section of the side bar. This will let you add molecules that are opened. (E.g. from SDF, Mol2, or mmCIF files), and set which atom indices in the trajectory they correspond to.