Molecule metadata
You can view metadata for a given molecule by clicking the Metadata button in the UI. For example, this will list data about, in the case of proteins, how it was imaged using crystallography, Cryo-EM etc. You can find more information using the database buttons, e.g. ones labeled PDBe, PubChem, DrugBank, RSCB etc. Which of these are present, and what metadata is available depends on molecule type, and information provided in its source file. Generally Metadata is robust for protein files from RCSB PDB, but may be sparse or non-existent for small molecules.
If you are loading or creating a novel molecule, you will have to populate this yourself!
Information may include the following: - PubChem, DrugBank, and other database IDs, - SMILES - InCHi - InChiKey - IUPAC name - Common name - Partial charge data - Atom count
Small molecule characterization
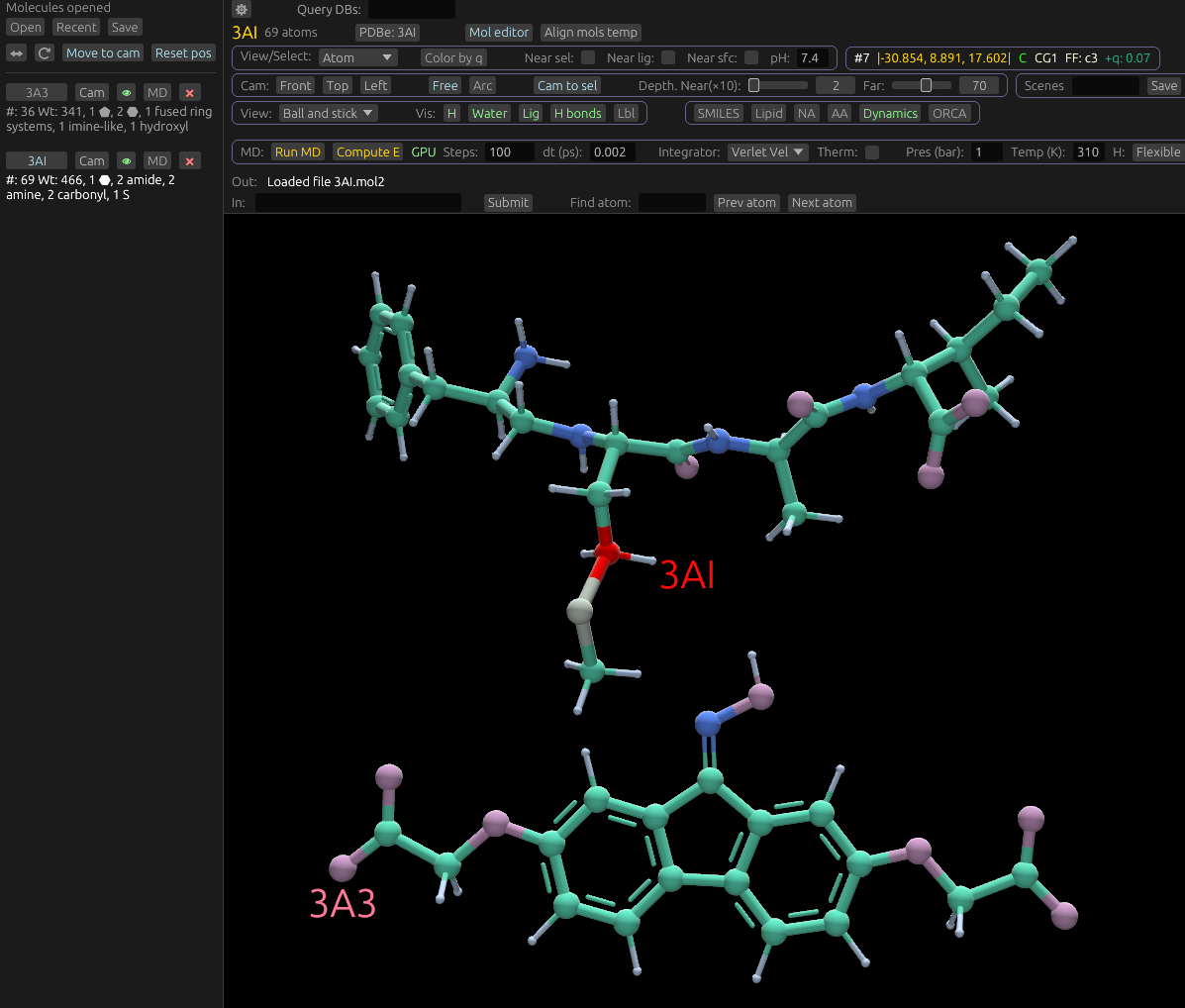
When Molchanica opens a small organic molecule, it characterizes its broad features, and displays them under its entry in the sidebar. See below for an example displaying two molecules, and the features it's characterized:
Some things it can list, computed analtically
- Atom count
- Molecular weight
- Rings, broken down symbolically by atom count and aromaticity
- Fused ring systems; these are counted independently of individual rings.
- Amine groups
- Amide groups
- Imine-like, Pyrrole-like, and pyridine-line nitrogens
- Carbonyl groups
- hydroxyl groups
- Less-common atoms like Sulfur, Phosphorus, and Chlorine
- Halogen count
- Number of rotatable bonds
- Hydrogen bond donors and acceptors
- LogP
- Molar refractivity
- Polar surface area
- Wiener index
We also compute and display pharmaceutical properties (e.g. ADME). For information on those, see the Therapeutic Properties section of this documentation.